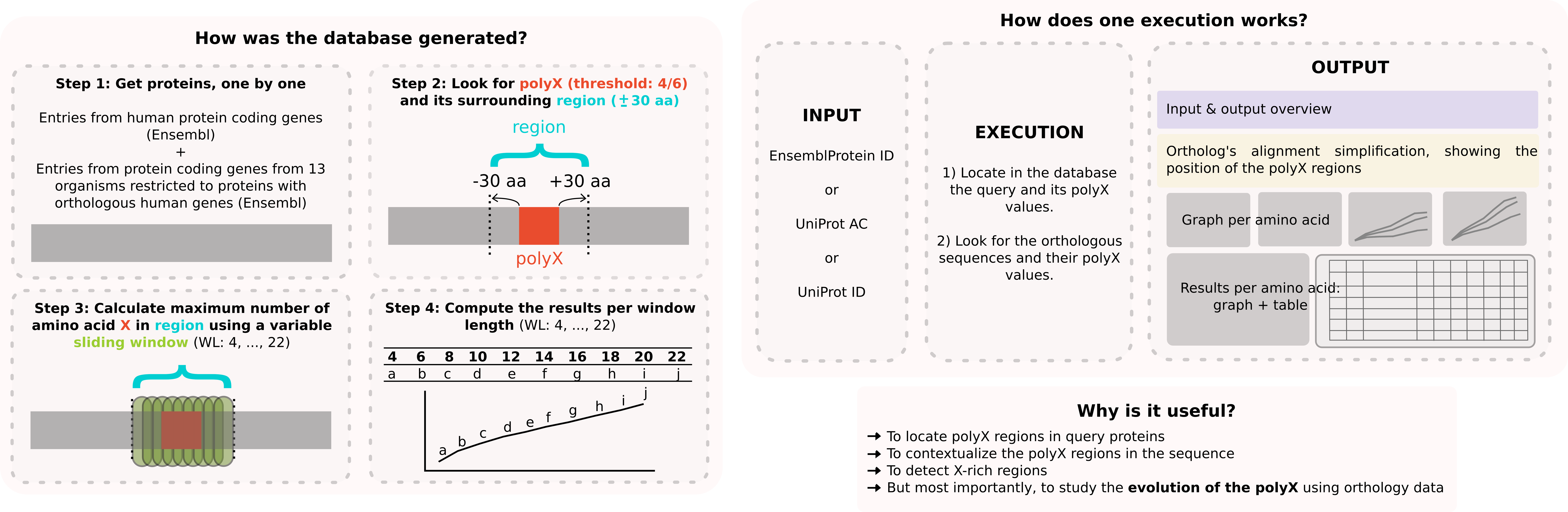
[Input option A] EnsemblProtein ID , UniProt AC or UniProt ID to get its polyX and their evolution using orthology data.
(Organisms in our database) *Execution time depends solely on the query length, from one second (~500 amino acids) to around 40 seconds (~4000 amino acids).
Homo sapiens (GRCh38.p7, 102147 entries)Pan troglodytes (CHIMP2.1.4, 18550 entries)Pongo abelii (PPYG2, 18524 entries)Macaca mulatta (MMUL_1, 31625 entries)Mus musculus (GRCm38.p4, 52335 entries)Rattus norvegicus (Rnor_6.0, 25005 entries)Bos taurus (UMD3.1, 20014 entries)Gallus gallus (Galgal4, 14202 entries)Xenopus tropicalis (JGI4.2, 17990 entries)Danio rerio (GRCz10, 32655 entries)Takifugu rubripes (FUGU4.0, 41397 entries)Ciona intestinalis (KH, 8368 entries)Caenorhabditis elegans (WBcel235, 12712 entries)Drosophila melanogaster (BDGP6, 18624 entries)
*Entries with orthologous human genes
Example 1 → one Ensembl ID (ENSP00000244769 , human ataxin 1) as query. Precomputed result.Example 2 → one UniProt AC (P42858 , human huntingtin) as query. Precomputed result.
[Input option B] protein sequence/s fasta format *Only the first 20 sequences will be computed and examined for polyX regions.  dAtabase of Polyx Evolution
dAtabase of Polyx Evolution dAtabase of Polyx Evolution
dAtabase of Polyx Evolution